Mapping Pandemics at the Library
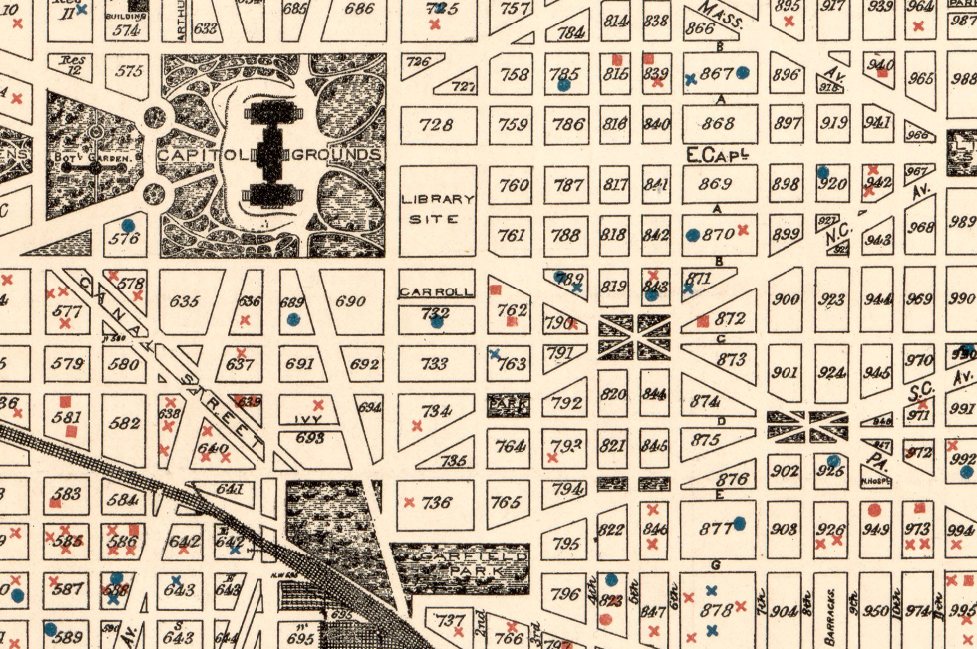
Sewer engineer’s map of Washington, D.C., 1894. Highlights show fatal cases of zymotic (infectious) diseases. Geography and Maps Division.
John Hessler is a specialist in the Library’s Geography and Map Division, focusing on computational geography and geographic information science. He’s also the Library’s curator of the Jay I. Kislak Collection. As part of our National Book Festival Presents series, he’ll be in conversation with Marie Arana (the Library’s literary director) to discuss the sweep of history from the 1500s smallpox pandemic that decimated the indigenous population of the Americas to the meticulous work that is being done now to map COVID-19. Please join us to watch this conversation tonight on the Library’s Facebook page. You can also watch it on our YouTube page.
John wrote this piece as a prologue to tonight’s conversation.
“Everyone knows that pestilences have a way of recurring in the world; yet somehow we find it hard to believe in ones that crash down on our heads from a blue sky.” Albert Camus, “The Plague.”
For the past month I have been involved in mapping the COVID-19 pandemic and searching for geospatial data and cartographic visualizations that will be important additions to the Library’s vast map collections. Future generations will rely on maps to help them understand this historical moment.
The mapping of infectious diseases and viral pathogens is nothing new; it became a scientific endeavor in the late 19th century. The most modern mapping technologies and statistical calculations were used to track viral and bacterial infections such as typhoid, malaria, scarlet fever, diphtheria and measles, which at the time were called zymotic (infectious) diseases. 
These infections were the subject of intense studies and mapping campaigns in large cities such as Washington, D.C. The studies were often carried out block-by-block, in order to determine the spread of pathogens and to plan for medical interventions.
This kind of historic high-scale mapping is important to epidemiologists of today, who study the past patterns of infectious disease transmission in order to understand the kind of pandemic we are now experiencing.
Today, our mapping technologies and fundamental understanding of the genomics of infectious disease are better than they were just a few decades ago. Several new data sources and providers have been organized in the last few years to provide information on the genomics and the spatial and temporal distribution of rapidly transmitted diseases in real time during outbreaks. For example, GISAID, the Global Initiative for Sharing All Influenza Data, aggregates genomic data from labs around the world during serious disease pandemics. They make that data available online.
That data is being used to map the outbreak of COVID-19. This has been the focus of much of what I have been adding to the Geography and Map Division collections. Combining the genomic data on the complex nucleotide mutations from these labs and then mapping how these mutations move around the globe has been a critical to how governments have responded to the outbreak.

Temporal phylogenetic tree of COVID-19. The purple represents the early form of the virus in China. The two red sections represent spread of the virus to North America from China and Europe. Courtesy Next Strai
This genomic data can be mapped to show how various transmission networks developed over the course of the pandemic. Preserving data and maps like this will be central to any future understanding of what is happening today.

Map of COVID-19 phylodynamics, Courtesy: Next Strain.
Like the high-scale mapping from the 19th century in our collections, it is important for us to collect geo-spatial data and maps like those mentioned here, and many others, all of which will allow future historians to study not only the spread of viral pathogens like COVID-19, but also help them comprehend how we as a culture used the best technologies available to us to react to this pivotal moment.
Subscribe to the blog— it’s free! — and the largest library in world history will send cool stories straight to your inbox.


Seja o primeiro a comentar